深度学习十二:PCA和whitening在二自然图像中的练习
前言:
现在来用PCA,PCA Whitening对自然图像进行处理。而本次试验的数据,步骤,要求等参考网页:http://deeplearning.stanford.edu/wiki/index.php/UFLDL_Tutorial 。实验数据是从自然图像中随机选取10000个12*12的patch,然后对这些patch进行99%的方差保留的PCA计算,最后对这些patch做PCA Whitening和ZCA Whitening,并进行比较。
实验环境:matlab2012a
实验过程及结果:
随机选取10000个patch,并显示其中204个patch,如下图所示:
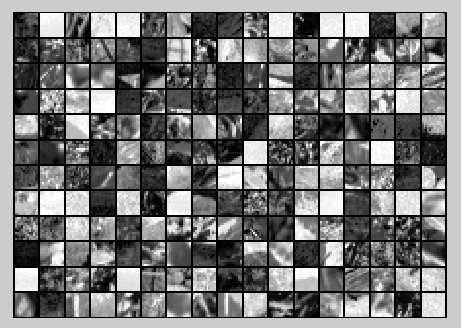

然后对这些patch做均值为0化操作得到如下图:

对选取出的patch做PCA变换得到新的样本数据,其新样本数据的协方差矩阵如下图所示:

保留99%的方差后的PCA还原原始数据,如下所示:

PCA Whitening后的图像如下:
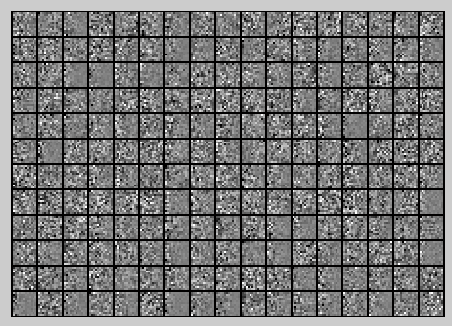

此时样本patch的协方差矩阵如下:
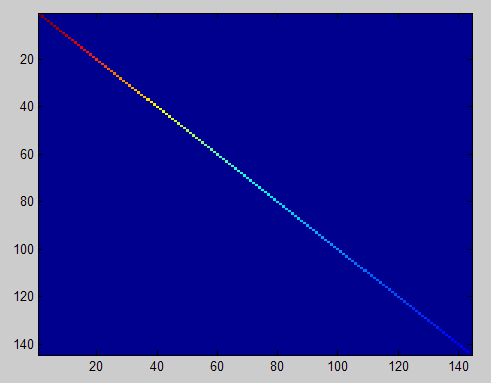
ZCA Whitening的结果如下:
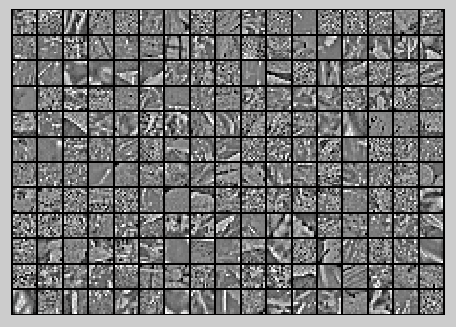

实验代码及注释:
%%================================================================
%% Step 0a: Load data
% Here we provide the code to load natural image data into x.
% x will be a 144 * 10000 matrix, where the kth column x(:, k) corresponds to
% the raw image data from the kth 12x12 image patch sampled.
% You do not need to change the code below.
x = sampleIMAGESRAW();
figure('name','Raw images');
randsel = randi(size(x,2),204,1); % A random selection of samples for visualization
display_network(x(:,randsel));%为什么x有负数还可以显示?
%%================================================================
%% Step 0b: Zero-mean the data (by row)
% You can make use of the mean and repmat/bsxfun functions.
% -------------------- YOUR CODE HERE --------------------
x = x-repmat(mean(x,1),size(x,1),1);%求的是每一列的均值
%x = x-repmat(mean(x,2),1,size(x,2));
%%================================================================
%% Step 1a: Implement PCA to obtain xRot
% Implement PCA to obtain xRot, the matrix in which the data is expressed
% with respect to the eigenbasis of sigma, which is the matrix U.
% -------------------- YOUR CODE HERE --------------------
xRot = zeros(size(x)); % You need to compute this
[n m] = size(x);
sigma = (1.0/m)*x*x';
[u s v] = svd(sigma);
xRot = u'*x;
%%================================================================
%% Step 1b: Check your implementation of PCA
% The covariance matrix for the data expressed with respect to the basis U
% should be a diagonal matrix with non-zero entries only along the main
% diagonal. We will verify this here.
% Write code to compute the covariance matrix, covar.
% When visualised as an image, you should see a straight line across the
% diagonal (non-zero entries) against a blue background (zero entries).
% -------------------- YOUR CODE HERE --------------------
covar = zeros(size(x, 1)); % You need to compute this
covar = (1./m)*xRot*xRot';
% Visualise the covariance matrix. You should see a line across the
% diagonal against a blue background.
figure('name','Visualisation of covariance matrix');
imagesc(covar);
%%================================================================
%% Step 2: Find k, the number of components to retain
% Write code to determine k, the number of components to retain in order
% to retain at least 99% of the variance.
% -------------------- YOUR CODE HERE --------------------
k = 0; % Set k accordingly
ss = diag(s);
% for k=1:m
% if sum(s(1:k))./sum(ss) < 0.99
% continue;
% end
%其中cumsum(ss)求出的是一个累积向量,也就是说ss向量值的累加值
%并且(cumsum(ss)/sum(ss))<=0.99是一个向量,值为0或者1的向量,为1表示满足那个条件
k = length(ss((cumsum(ss)/sum(ss))<=0.99));
%%================================================================
%% Step 3: Implement PCA with dimension reduction
% Now that you have found k, you can reduce the dimension of the data by
% discarding the remaining dimensions. In this way, you can represent the
% data in k dimensions instead of the original 144, which will save you
% computational time when running learning algorithms on the reduced
% representation.
%
% Following the dimension reduction, invert the PCA transformation to produce
% the matrix xHat, the dimension-reduced data with respect to the original basis.
% Visualise the data and compare it to the raw data. You will observe that
% there is little loss due to throwing away the principal components that
% correspond to dimensions with low variation.
% -------------------- YOUR CODE HERE --------------------
xHat = zeros(size(x)); % You need to compute this
xHat = u*[u(:,1:k)'*x;zeros(n-k,m)];
% Visualise the data, and compare it to the raw data
% You should observe that the raw and processed data are of comparable quality.
% For comparison, you may wish to generate a PCA reduced image which
% retains only 90% of the variance.
figure('name',['PCA processed images ',sprintf('(%d / %d dimensions)', k, size(x, 1)),'']);
display_network(xHat(:,randsel));
figure('name','Raw images');
display_network(x(:,randsel));
%%================================================================
%% Step 4a: Implement PCA with whitening and regularisation
% Implement PCA with whitening and regularisation to produce the matrix
% xPCAWhite.
epsilon = 0.1;
xPCAWhite = zeros(size(x));
% -------------------- YOUR CODE HERE --------------------
xPCAWhite = diag(1./sqrt(diag(s)+epsilon))*u'*x;
figure('name','PCA whitened images');
display_network(xPCAWhite(:,randsel));
%%================================================================
%% Step 4b: Check your implementation of PCA whitening
% Check your implementation of PCA whitening with and without regularisation.
% PCA whitening without regularisation results a covariance matrix
% that is equal to the identity matrix. PCA whitening with regularisation
% results in a covariance matrix with diagonal entries starting close to
% 1 and gradually becoming smaller. We will verify these properties here.
% Write code to compute the covariance matrix, covar.
%
% Without regularisation (set epsilon to 0 or close to 0),
% when visualised as an image, you should see a red line across the
% diagonal (one entries) against a blue background (zero entries).
% With regularisation, you should see a red line that slowly turns
% blue across the diagonal, corresponding to the one entries slowly
% becoming smaller.
% -------------------- YOUR CODE HERE --------------------
covar = (1./m)*xPCAWhite*xPCAWhite';
% Visualise the covariance matrix. You should see a red line across the
% diagonal against a blue background.
figure('name','Visualisation of covariance matrix');
imagesc(covar);
%%================================================================
%% Step 5: Implement ZCA whitening
% Now implement ZCA whitening to produce the matrix xZCAWhite.
% Visualise the data and compare it to the raw data. You should observe
% that whitening results in, among other things, enhanced edges.
xZCAWhite = zeros(size(x));
% -------------------- YOUR CODE HERE --------------------
xZCAWhite = u*xPCAWhite;
% Visualise the data, and compare it to the raw data.
% You should observe that the whitened images have enhanced edges.
figure('name','ZCA whitened images');
display_network(xZCAWhite(:,randsel));
figure('name','Raw images');
display_network(x(:,randsel));参考资料:
Deep learning:十(PCA和whitening)
http://deeplearning.stanford.edu/wiki/index.php/UFLDL_Tutorial
作者:tornadomeet
出处:http://www.cnblogs.com/tornadomeet 欢迎转载或分享,但请务必声明文章出处。
新浪微博:tornadomeet,欢迎交流!
下一篇:深度学习十一:PCA和whitening在二维数据中的练习
[按键盘← → 左右方向键也可以翻页哦~]
[按键盘← → 左右方向键也可以翻页哦~]






